The company recently attended a J.P. Morgan Healthcare Conference in San Francisco last month (January 7-10), where it attracted interest from investors in its technology including Inflammatix molecular diagnostics company, developing tests that read the immune system.
BARDA grant

“Our technology is a genetic analyzer but we can do more than pathogen detection, we can quantify expression levels of genes, whether they are part of a bacterial response or the host response, ie if a person is infected their immune cells respond to the infection and this triggers a cascade of events at higher levels,” said Jack Regan, CEO, LexaGene.
“Take sepsis for example, sepsis that gets into your blood deregulates the immune system and the way you measure it is by certain ‘markers’ that go out of whack, it’s important to quantify this.
“We are a public company, we wanted to talk to the owner of Inflammatix at the conference and we will submit some grants together, we want to apply for a BARDA grant, part of the Biomedical Advanced Research and Development Authority. They fund a lot of startups and they are interested in solving things like sepsis, which kills up to 270,000 people a year in the US and the amount of money dealing with that is extremely high, also influenza.”
LexeGene and Inflammatix hope to hear if the grant has been accepted in about two months’ time. Traditionally, Regan said the turnaround for applications used to be around nine months but the BARDA DRIVe Program mandate is to accelerate the development of new technology to help impact areas.
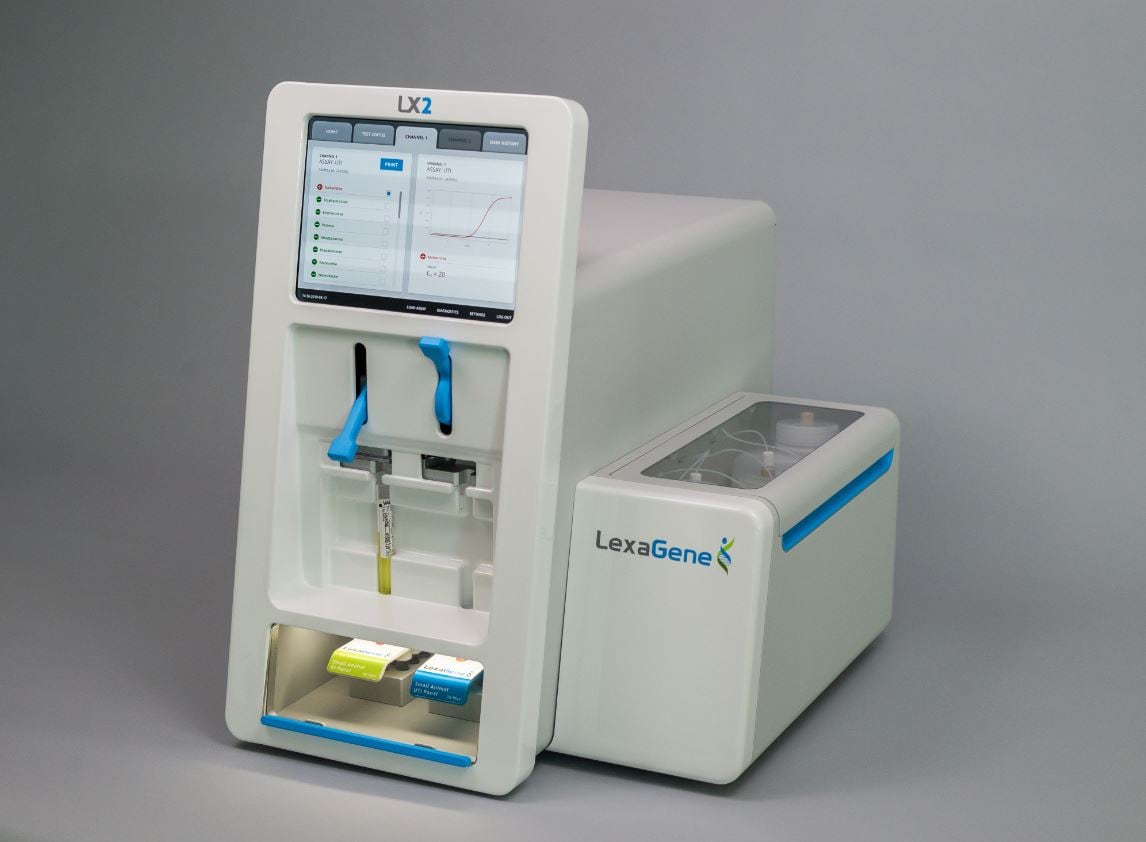
Regan says LexaGene is currently in the process of developing LX2, ‘the world’s first on-site, open-access instrumentation for pathogen detection’ to be used for food safety, veterinary diagnostics and water quality testing. Open-access means it can be customized for up to 22 types of genetic screening.
He said it is important to note it is the first ever ‘open-access’ technology because others are ‘closed –access’ where the vendor sends the test to a third party and tells the client what to look for, which can be done at a high cost.
“In human clinical diagnostics the food sector is different. You screen to detect certain pathogens. With our company we are low cost, open-access, and we are designing and validating assets for food safety so they don’t have to send it to a lab,” added Regan.
“We are confident we will have a lot of traction. We already have interest from one of the largest producers of organic pre-made salads who want to work with us.”
Regan said LexeGene is also in talks with an attorney to figure out if food companies would be subject to increased liability or not using their testing methods.
“A lot of food companies are concerned about liability right now. When they do food testing they send their samples to a third party vendor, who is certified and runs AOAC tests, or AFNOR tests if it’s in Europe. The question is if they evaluate our technology and it comes up with a positive result. Traditionally tests are negative, so although we are not yet certified we want to know if they would be liable to a greater extent should that food product be clear to go.
“We are going through an attorney to figure out if they (the food companies) would be subject to increased liability as we don’t want them to have more risk. We will get certification but at the moment we are still in product development mode.”
Commercially available 2019
Following tests LexeGene is hoping to have completed data testing on its technology by summer and submit a certificate to AOAC to go on the market by the end of this year, for companies to adopt its technology and to be compliant with the regulations.
“We are not targeting restaurants but suppliers, packaging manufacturers and processors of food. Anyone who is touching food, for multi-state distribution, food contract labs, Government food labs and big retail chains such as Walmart and Costco, who do their own testing or ask their suppliers to do it,” added Regan.
“We are targeting all these people and we are trying to be the first to provide broad-spectrum genetic testing to the food industry. There is such a demand for low-cost testing.
“The industry often selects one or two pathogens to test and turns a blind eye to the others but there are 12 major pathogens that cause major food illnesses. We want to provide a broad-spectrum screening so they feel more comfortable with their food testing procedures to ensure their livelihood is not at stake with any recalls.”
With its LX2 genetic analyzer the company claims it can provide results in one hour – where current methods return results in one to three days – and says it can test up to six samples at a time with a high degree of specification.
The way the technology works is by taking a raw sample and evaluating its genetics via a liquid. The platform is easy-to-use: end-users collect a sample, load it onto the instrument with a sample preparation cartridge, and press ‘go’. The instrument is expected to offer excellent sensitivity, specificity, and breadth of pathogen detection—with the ability to provide test results in one hour – as opposed to three to five days with current methods – and can screen up to 22 pathogens at once.
“We can’t process chunks of meat or products such as peanut butter. It has to be in liquid form and not too viscous,” said Regan.
“In our case the liquid would be drawn up into a single usage cartridge, that is designed to concentrate the pathogens inside the sample, then purify the genetic material form to detect for pathogens.
“The concentration step is very important, the purified genetic material enters the instrument, and the tests, based on PCR (Polymerase Chain Reaction), can screen up to 28 targets at once. The results are quantitative.”
Salmonella and E.coli
Regan added the desire to confirm viability - traditionally the technology that has been used has been based on culture testing - genetics have the advantage of being quick and sensitive but can’t confirm viability but in a shorter timeframe it can measure the amount of DNA and the sample can be measured six hours after.
Current testing methods in the food safety industry use culture-based methods to ensure a product is safe for consumption before delivering to a store.
This method of testing involves taking a sample of the product and measuring the growth of bacteria under controlled conditions (added to an enriched broth, and incubated at 37C under aerobic conditions) over a period of time. These samples require a minimum of 24-48 hours of growth before being analyzed and returning a result, which means food processors must wait several days to confirm if their products are safe for consumption.
“If the amount of pathogen material goes up that suggests bacteria is viable and growing, and from a food safety perspective, a lot of companies do not have a kill step in their processing for food such as salads, so they are at high risk of contamination and it’s a perishable product,” he said.
“The shelf life of a salad is about 10 days so if their product is in storage for two days while they are waiting for viability tests to come back this shortens the timeframe to sell their product. Right now the processing time is 24 hours for something like listeria (12 hours for salmonella and E.coli) for example so they can’t ship it out any faster than that.
“Listeria takes longer to replicate but with our technology it’s so sensitive you can take a sample on time point zero and the anticipation is you look at it six hours later to retest it.
“Our goal is to test the sample twice at low cost and clear it ready to be shipped within eight hours which is a big deal in comparison to waiting 12 hours.”
LexaGene hopes to ship its beta technology to customers by next summer, and the technology is slated for commercialization in 2019. The company plans to ship its beta technologies to customers by summer, and the technology is slated for commercialization in 2019.
The implications of this technology for the food industry is huge says Regan - foodborne illnesses cost the US economy between $55 and $93 billion annually and the CDC estimates 48 million people get sick from foodborne diseases each year in the US.
Similarly, the water quality testing market is expected to reach $3.5 billion by 2019, with testing needs for municipal water systems, water waste treatment plants, sewage overflow, and beaches.
Regan completed his doctoral training at the University of California, San Francisco (UCSF), where he studied influenza, and performed his post-doctoral studies at Lawrence Livermore National Laboratories, where he was a lead scientist in developing the APDS instrument that was adopted by the Department of Homeland Security (DHS) for use in its BioWatch program for continuous biothreat surveillance.
He previously worked for biotech companies Applied Biosystems, Life Technologies, Bio-Rad Laboratories and QuantaLife, where he worked on automated sample preparation and developed/commercialized TaqMan-based reagents to detect pathogens, cancer and neurological disorders.
“Through these experiences I fell into this career path making it more sensitive and easier to detect pathogens and designing new instruments,” he added.
Highlighting the fundamental issues he believes the food industry needs to address immediately to prevent people from getting sick from widespread foodborne illnesses, Regan said firms need to adopt genetic testing to get a risk assessment to get results faster because currently the culture timing is too slow.
All pathogens
“We need to adopt genetic testing to allow for a broader screening looking at all pathogens regarding food safety,” he said.
Time point 0 means the moment the sample is added to the broth, an aliquot (small volume) is removed for testing. Time point = six hours would mean after six hours of ‘controlled conditions,’another small aliquot is removed and compared to the data generated from Time point = 0.
“The problem with culture is it doesn’t look for everything ie the Norovirus. One of the concerns right now is genetics is so sensitive it not only tests dead but alive pathogens. A lot of what the industry does is kill step false positives, but genetics can get a product in and out of the door within eight instead of 24 hours.
“If you do one time point you can assess the risk if there is DNA from a pathogen but we don’t know if it’s alive or dead.
“If there is nothing there you know you are free to ship it. Negative means it’s cultured but maybe positive after 24 hours. The whole idea is our technology can rapidly assess risk. If there is no risk it can be shipped right away.
“If there is risk detect viability and how the customer decides to use that data is up to them. This is a new technology to the field so there is going to be a growing period how food genetics fits into the food industry and there will be some challenges but we feel it is low cost and the food industry is in desperate need of innovation and this is a great solution for them.”
According to Regan the most at-risk food types are salad, leafy greens, chicken and beef. Meat is always traditionally the highest risk for E.coli and salmonella, and they have a lot of kill steps.
“We are looking to provide an extra layer of security in pathogen testing. The minute you detect that pathogen extreme you need to be on high alert as to how that food was processed what machinery was used and where was it packaged,” added Regan.
“The amount of DNA in meat is much higher than in fruit and vegetables that has gone through three washes. There is also the environment where bacteria can grow on raw meat.
For example, Arizona meat producer JBS Tolleson expanded a ground beef meat recall to include more than 12 million pounds of product that was packaged from July to September 2018, and sold under the Showcase, Cedar Farms, and Kroger brands.
The USDA’s Food Safety and Inspection Service believed the ground beef was contaminated with a strain of salmonella. A study showed 246 people in 26 states were sick from the beef since the summer.
Last year the US CDC (Centers for Disease Control & Prevention), public health and regulatory officials in several states, CanadaExternal, and the US Food and Drug Administration (FDA) investigated a multi-state outbreak of Shiga toxin-producing Escherichia coli O157:H7 (E. coli O157:H7) infections linked to romaine lettuce from the Central Coastal growing regions in northern and central California.
“With the ground beef and romaine lettuce recalls last year alone, it shows the current technology isn’t working well enough. Here at LexeGene we don’t expect to replace the technology for testing but to compliment it for more rapid information,” added Regan.
LexeGene based north of Boston currently has 21 employees and is hoping to grow its team in the next couple of months. It will be attending IAFP from July 21-24 at Kentucky International Convention Center in Louisville, Kentucky.